Introduction
What are African trypanosomes and why do we study them?
African trypanosomes are parasites, spread between mammals by tsetse flies. They are responsible for epidemics of sleeping sickness in sub-Saharan Africa. This disease is the second biggest killer behind HIV in parts of Africa, but probably has more important impact through its effects on livestock, this leading to significant economic hardship and restricted development in afflicted regions. These parasites swim in our blood but can avoid being destroyed by antibodies raised against them by regularly changing the proteins on their surface. This process, antigenic variation, allows them to survive and multiply for long periods in the blood of humans, livestock and game animals in Africa. Unless treated, sleeping sickness is fatal; however current drugs are very old (mainly developed over 60 years ago) and have dangerous side effects.
Trypanosomes are very ancient and interesting organisms: they are among the earliest branching eukaryotes. Fundamental discoveries have been made in these parasites which have had major impact on what we know about all cells. They also are unique in many respects, an example of the “differently evolved”.
Trypanosomes as a tractable model for developmental processes.
We are interested in how trypanosomes control their life cycle, which governs the spread of disease. This is very accessible to molecular cell biology: different stages can be cultured in vitro, they can be genetically transformed, gene transcripts can be ablated by RNA interference and differentiation between life stages is simple and reproducible. Also the genome sequence of these parasites has now been completed, revealing that ~60% of trypanosome genes are not found in other organisms. It is these genes that may provide the keys to understanding the unique and interesting biology of trypanosomes.
The life cycle of trypanosomes
Trypanosomes survive free in the bloodstream of mammalian hosts. Here they proliferate as slender forms. However, as parasite density increases they differentiate to so-called stumpy forms using an inter-cellular communication system similar to quorum sensing observed in some microbial communities. Stumpy forms do not divide and are pre-adapted for survival and differentiation when taken up in the bloodmeal of a tsetse fly. Stumpy forms are pivotal in the transmission of trypanosomes, being more robust than slender forms and competent to differentiate. What defines the differentiation-competence of stumpy forms is one of our major research questions, and we have identified several key molecules that are important in this differentiation competence. One family of these are PAD proteins, responsible for helping the trypanosome to detect its uptake in a tsetse bloodmeal.
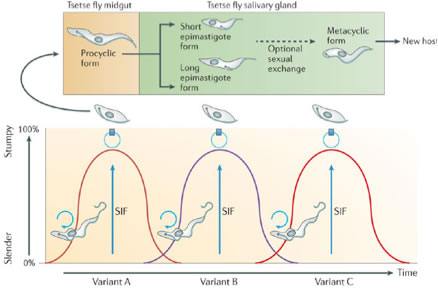
The life cycle of African Trypanosomes in the mammalian bloodstream and tsetse fly (from MacGregor, Szoor, Savill & Matthews;
Nature Reviews Microbiology 10, 431-438 (2012)
Once stumpy forms enter the tsetse fly midgut they differentiate into procyclic forms. This differentiation can be carried out in vitro and is completely synchronous. This is a great advantage for studies at the molecular and cellular level, because it enables the analyses of large numbers of parasites at the molecular level, and direct translation of this information into what is happening in individual cells. Using this synchronous differentiation model we have been able to map the major events of differentiation, with the aim to identify their underlying controls. How the programme of developmental events is controlled and co-ordinated is a second major research theme in our laboratory.
Stumpy forms isolated at peak parasitaemia
Understanding the life cycle control of trypanosomes has the potential to provide routes to preventing transmission of the disease. It also represents a model for eukaryotic cell differentiation in an organism that diverged far earlier than all other eukaryotic experimental models. This has the potential to identify conserved processes fundamental in eukaryotic cell biology. For example, cell type differentiation often involves progression of a proliferative cell (e.g. a stem cell) into cell-cycle arrest, this being followed by cell specialization via differentiation. The trypanosome life cycle follows a similar path: proliferative cells (bloodstream slender forms) undergo cell cycle arrest in response to an autocrine signal, after which the arrested cells (stumpy forms) differentiate in response to environmental cues to produce a new differentiated (but proliferative) cell type (procyclic cells).
A video highlighting our research work recognised by the Sanofi-Institut Pasteur prix for Tropical Medicine can be viewed here

